Solution. Interpretation of \(\gamma_0\):
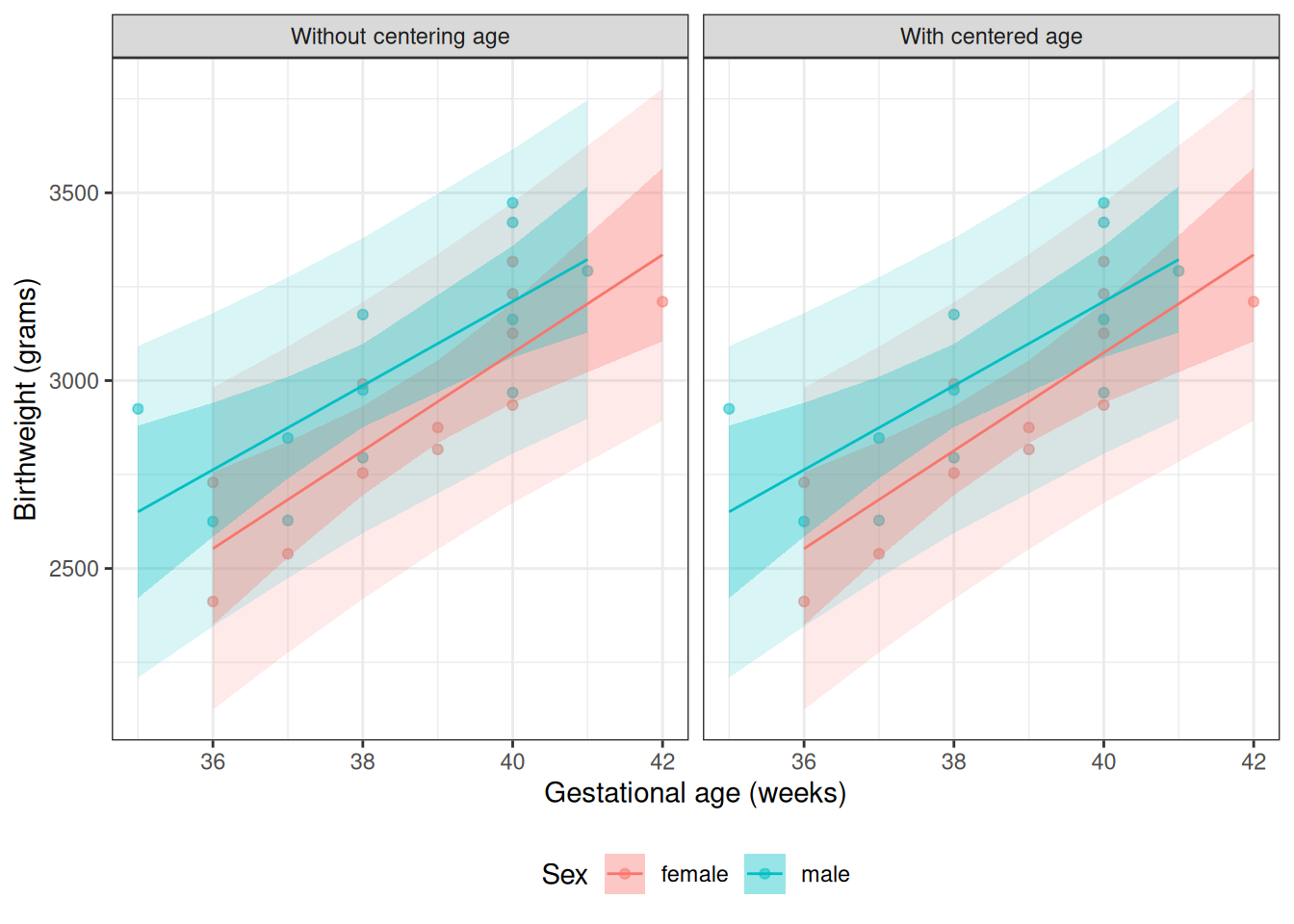
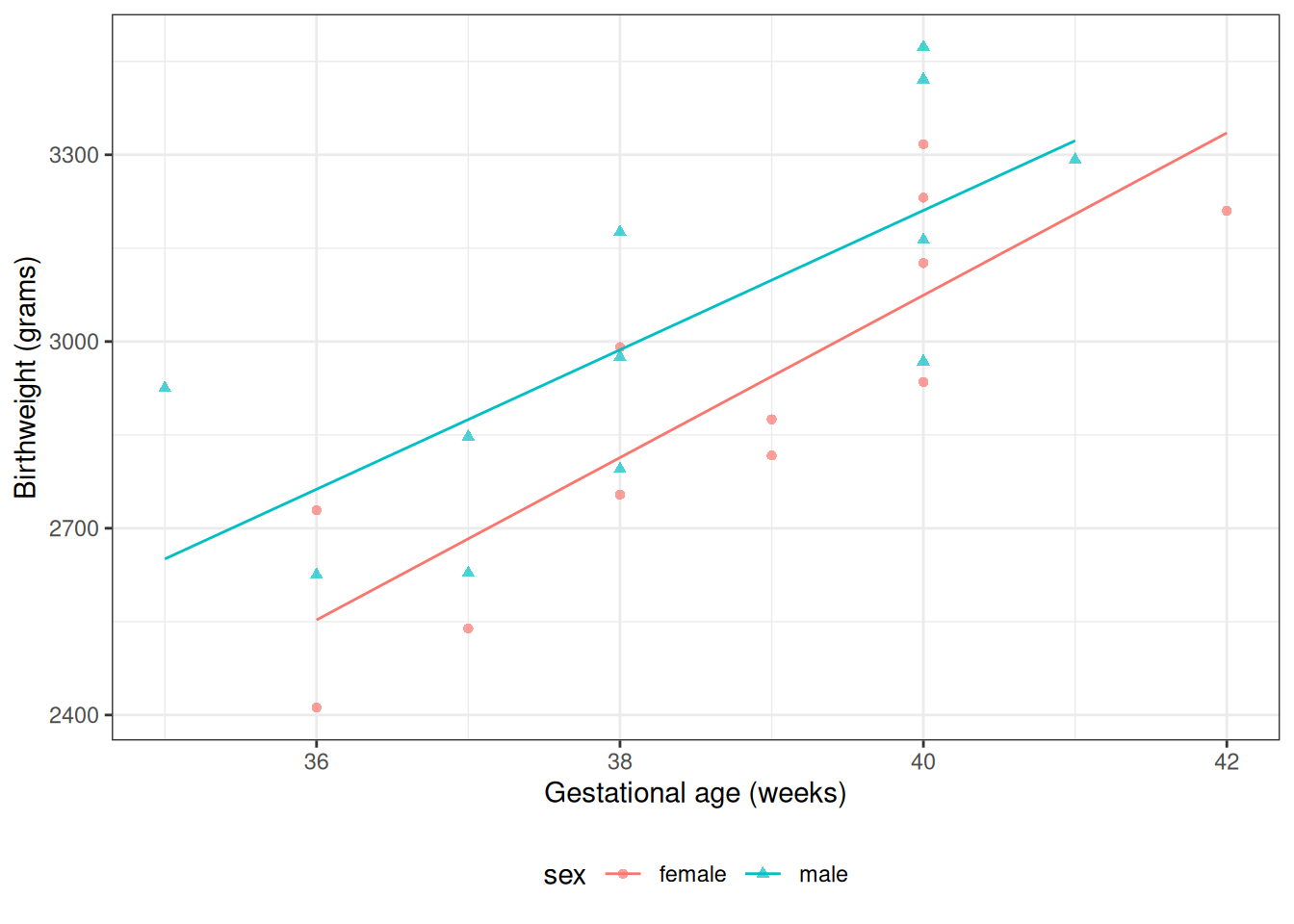
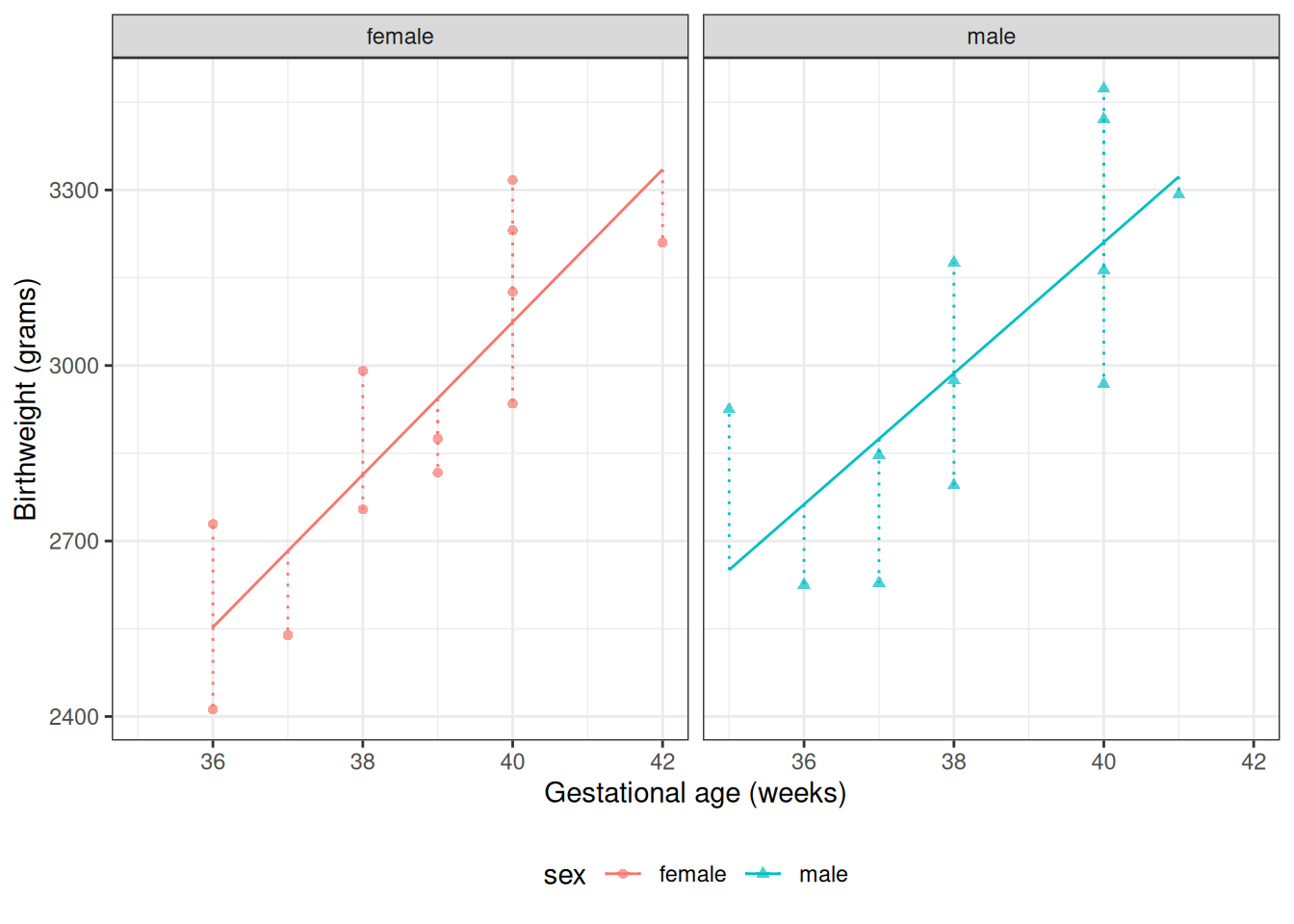
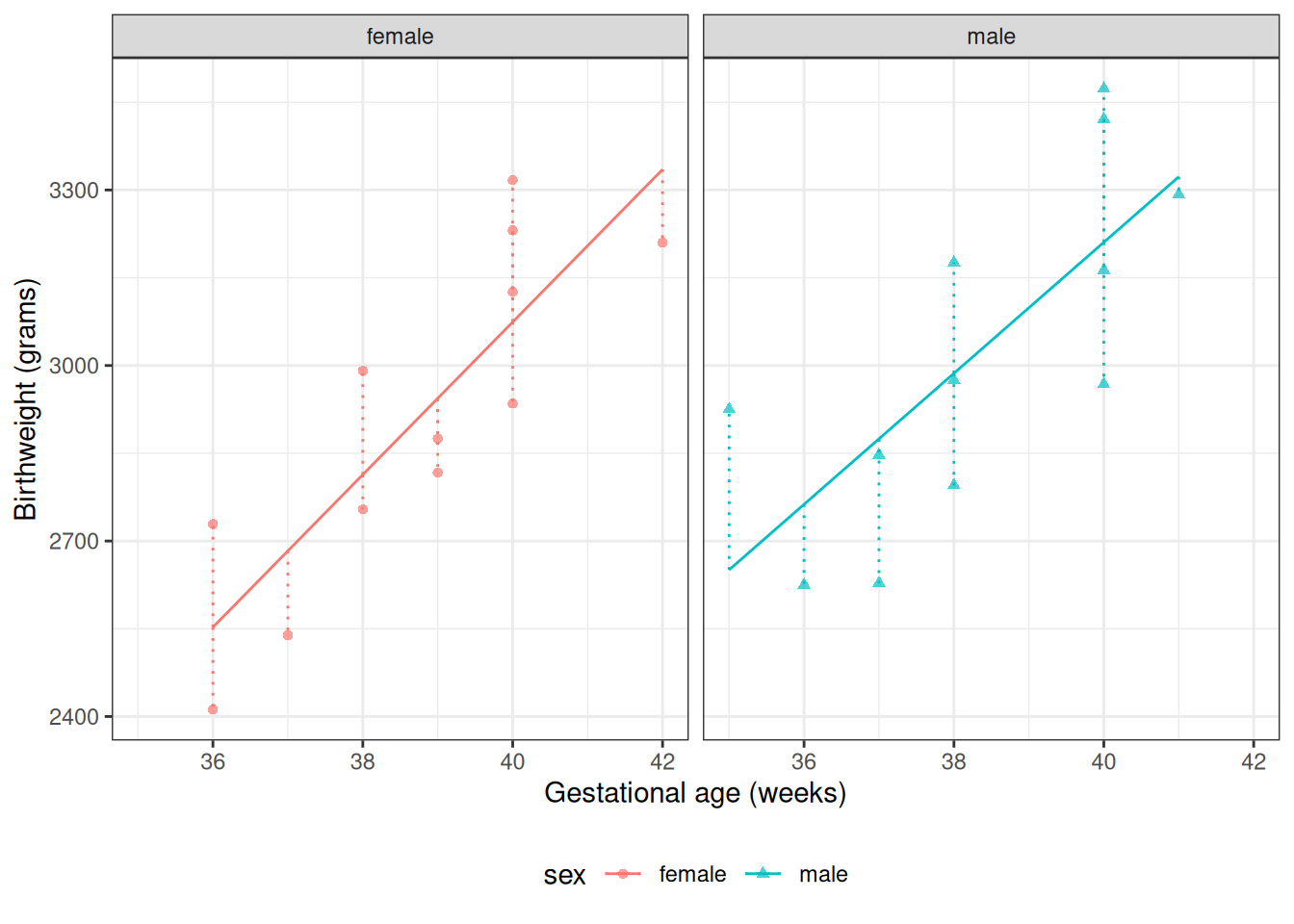
From Model 4, \(\gamma_0\) is the mean birthweight among females (\(M = 0\)) with \(A^* = 0\) (i.e., \(A = 32\) weeks):
\[
\gamma_0 = \text{E}{\left[Y|M=0, A^*=0\right]} = \text{E}{\left[Y|M=0, A=32\right]}
\]
Substituting into Model 2:
\[
\begin{aligned}
\gamma_0
&= \beta_0 + \beta_M \cdot 0 + \beta_A \cdot 32 + \beta_{AM} \cdot 0 \cdot 32\\
&= \beta_0 + 32\beta_A
\end{aligned}
\]
This differs from \(\beta_0 = \text{E}{\left[Y|M=0, A=0\right]}\), which is the mean birthweight among females at \(A = 0\) weeks.
Interpretation of \(\gamma_M\):
From Model 4, \(\gamma_M\) is the sex difference in mean birthweight at \(A^* = 0\) (i.e., \(A = 32\) weeks):
\[
\begin{aligned}
\gamma_M
&= \text{E}{\left[Y|M=1, A^*=0\right]} - \text{E}{\left[Y|M=0, A^*=0\right]}\\
&= \text{E}{\left[Y|M=1, A=32\right]} - \text{E}{\left[Y|M=0, A=32\right]}
\end{aligned}
\]
Substituting into Model 2:
\[
\begin{aligned}
\gamma_M
&= (\beta_0 + \beta_M + \beta_A \cdot 32 + \beta_{AM} \cdot 32) - (\beta_0 + \beta_A \cdot 32)\\
&= \beta_M + 32\beta_{AM}
\end{aligned}
\]
This differs from \(\beta_M\), which is the sex difference at \(A = 0\) weeks.
Interpretation of \(\gamma_{A^*}\):
From Model 4, \(\gamma_{A^*}\) is the slope of mean birthweight with respect to \(A^*\) among females (\(M = 0\)):
\[
\begin{aligned}
\gamma_{A^*}
&= \frac{d}{da^*}\text{E}{\left[Y|M=0, A^*=a^*\right]}\\
&= \frac{d}{da^*}\text{E}{\left[Y|M=0, A=a^*+32\right]}\\
&= \frac{d}{da}\text{E}{\left[Y|M=0, A=a\right]}
\end{aligned}
\]
Substituting into Model 2:
\[
\begin{aligned}
\gamma_{A^*}
&= \frac{d}{da}\left(\beta_0 + \beta_A \cdot a\right)\\
&= \beta_A
\end{aligned}
\]
Since shifting \(A\) by a constant does not change the slope, \(\gamma_{A^*} = \beta_A\): these two coefficients have the same value and interpretation.
Interpretation of \(\gamma_{A^*M}\):
From Model 4, \(\gamma_{A^*M}\) is the difference in slope with respect to \(A^*\) between males and females:
\[
\begin{aligned}
\gamma_{A^*M}
&= \frac{d}{da^*}\text{E}{\left[Y|M=1, A^*=a^*\right]} - \frac{d}{da^*}\text{E}{\left[Y|M=0, A^*=a^*\right]}\\
&= \frac{d}{da}\text{E}{\left[Y|M=1, A=a\right]} - \frac{d}{da}\text{E}{\left[Y|M=0, A=a\right]}
\end{aligned}
\]
Substituting into Model 2:
\[
\begin{aligned}
\gamma_{A^*M}
&= (\beta_A + \beta_{AM}) - \beta_A\\
&= \beta_{AM}
\end{aligned}
\]
Since shifting \(A\) by a constant does not change slopes, \(\gamma_{A^*M} = \beta_{AM}\): these two coefficients have the same value and interpretation.
The pattern:
Slope coefficients (\(\gamma_{A^*}\) and \(\gamma_{A^*M}\)) are unchanged by rescaling: they have the same values and interpretations as the corresponding \(\beta\)s.
Coefficients change only for variables that have interactions with the rescaled variable \(A\). This includes the intercept (which can be viewed as the main effect of a variable that interacts with \(A\) via \(\beta_A\)), and the main effect of \(M\) (which interacts with \(A\) via \(\beta_{AM}\)). Shifting \(A\) by 32 weeks changes the reference point from \(A = 0\) to \(A = 32\), so these coefficients now represent quantities evaluated at \(A = 32\) weeks rather than at \(A = 0\) weeks.